About the Journal
Scope of the Journal
Indexed In
Journal Information
Current issue
Archive
Editorial Board
Journal Policy
APC Policy
Authorship Policy
Author Recognition Policy
Citation Policy
Conflict of Interest (COI) Policy
Copyright Policy
Data Availability Policy
DEI Policy
Digital Archiving Policy
Ethical Approval Policy
Funding Policy
Image Policy
Open Access Policy
Ownership & Governance
Peer Review Policy
Personal Data Protection Policy
Plagiarism Policy
Post-Publication Policy
Preprint Policy
Author Instructions
Submit Manuscript
Contact Information
Journal Policy
APC Policy
Authorship Policy
Author Recognition Policy
Citation Policy
Conflict of Interest (COI) Policy
Copyright Policy
Data Availability Policy
DEI Policy
Digital Archiving Policy
Ethical Approval Policy
Funding Policy
Image Policy
Open Access Policy
Ownership & Governance
Peer Review Policy
Personal Data Protection Policy
Plagiarism Policy
Post-Publication Policy
Preprint Policy
RESEARCH PAPER
Computational Structural Analysis and Interaction Network Profiling of Lipases: Implications for Biomedical and Therapeutic Applications
1
Department of Crop Science, Faculty of Agriculture, Universiti Putra Malaysia, Malaysia
2
Department of Bioinformatics, Vels Institute of Science, Technology and Advanced Studies, India
Submission date: 2026-02-14
Final revision date: 2026-03-27
Acceptance date: 2026-04-01
Online publication date: 2026-04-28
Publication date: 2026-04-28
Corresponding author
Vasudevan Venkatachalam
Department of Crop Science, Faculty of Agriculture, Universiti Putra Malaysia, 43400, Selangor, Malaysia
Department of Crop Science, Faculty of Agriculture, Universiti Putra Malaysia, 43400, Selangor, Malaysia
Journal of Medico Informatics 2026;02(Issue 02):6-13
HIGHLIGHTS
- Lipases exhibit high catalytic versatility across aqueous and organic solvent environments.
- Structural stability depends on hydrophobic interactions, hydrogen bonds, and ionic interactions.
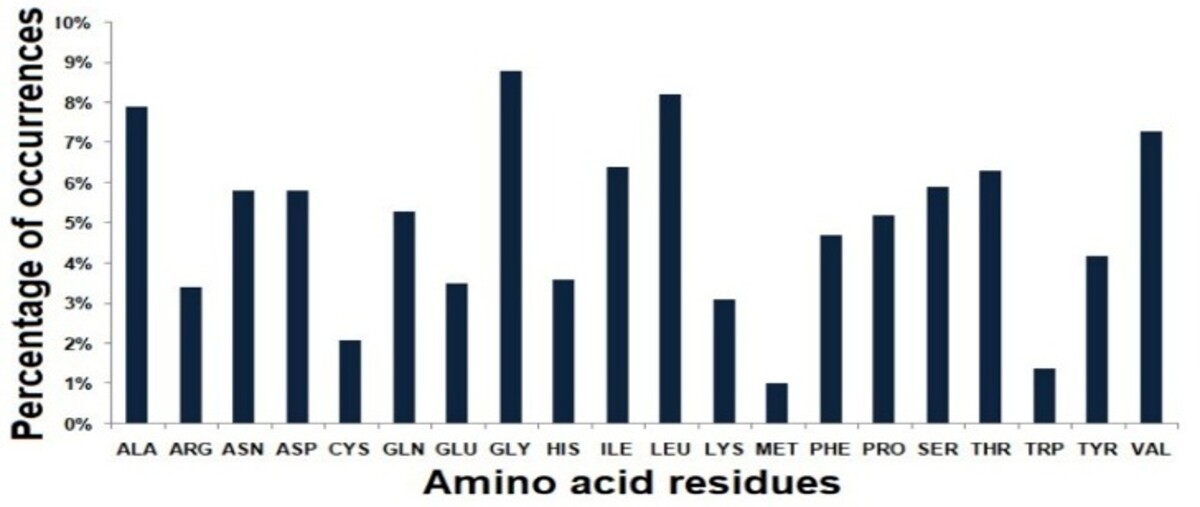
- Glycine-rich composition enhances flexibility and solvent tolerance in lipases
- Computational analysis identified key structural determinants of solvent-stable lipases.
- Protein–protein interaction networks reveal species-specific functional differences in lipases.
KEYWORDS
TOPICS
ABSTRACT
Lipases are widely used enzymes that facilitate the process of breaking down triglycerides to glycerol and fatty acids and are involved in different processes of lipid metabolism, drug metabolism, the diagnosis of diseases, and several industrial processes. In this paper, a detailed computational work has been conducted to examine the structural determinants of the stability of organic solvents in lipases. An eighty two lipase dataset was obtained in the Protein Data Bank and assessed by amino acid composition profiling, stabilizing structural interactions, and calculation of physicochemical properties. The parameters such as the hydrophobic interactions, salt bridges, hydrogen bonds, and packing density were determined to determine factors that contributed to the stability of the solvents. The analysis of protein-protein interaction with the STRING database also revealed the functional association of the lipases in the different species (including Homo sapiens, Pan troglodytes, and Mus musculus). This study has limitations, however, in terms of its being based on computational analysis and the small number of experimentally characterized solvent-stable lipases that can be compared. Thus, experimental validation by mutagenesis, soluble stability experiments with enzymes, and rational protein engineering strategies should be undertaken in future efforts to improve solvent tolerance level depending on the structural characteristics observed. The results can inform the structural aspects of lipase stability and can be used in the future in the fields of biomedical studies, drug development, and enzyme engineering.
ABBREVIATIONS
PDB – Protein Data Bank
PPI – Protein–Protein Interaction
ASA – Accessible Surface Area
VLDP – Voronoi Library of Domain Proteins
CHARMM – Chemistry at HARvard Macromolecular Mechanics
Å – Angstrom
pI – Isoelectric Point
MW – Molecular Weight
STRING – Search Tool for the Retrieval of Interacting Genes/Proteins
HEPES – 4-(2-hydroxyethyl)-1-piperazineethanesulfonic acid
Tris – Tris(hydroxymethyl)aminomethane
PEG – Polyethylene Glycol
DGAT – Diacylglycerol O-Acyltransferase
MGLL – Monoglyceride Lipase
FAAH – Fatty Acid Amide Hydrolase
ACKNOWLEDGEMENTS
The authors gratefully acknowledge Universiti Putra Malaysia for providing the necessary research infrastructure, computational resources, and institutional support required to carry out this research work. The authors also appreciate the conducive academic environment and technical assistance that facilitated the successful completion of this work.
FUNDING
This research received no external funding. All work was conducted using institutional resources without dedicated grant support.
CONFLICT OF INTEREST
The authors declare that they have no known financial, personal, academic, or other relationships that could inappropriately influence, or be perceived to influence, the work reported in this manuscript. All authors confirm that there are no competing interests to declare.
PEER REVIEW INFORMATION
Article has been screened for originality
© 2026 The Author(s). This article is distributed under the terms of the Creative Commons Attribution License (CC BY 4.0).
REFERENCES (54)
1.
Ait Lahcen N, Liman W, Zekri S, Ait Lahcen A, Alanazi AS, Alanazi MM, Delaite C, Maatallah M, Cherqaoui D. (2026), QSAR-Guided and Fragment-Based Drug Design of Monoterpenoid Inhibitors Targeting Ebola Virus Glycoprotein, Int J Mol Sci, 27(7) doi:10.3390/ijms27072987. PMID: 41977174.
2.
Ali S, Khan SA, Hamayun M, Lee IJ. (2023), The Recent Advances in the Utility of Microbial Lipases: A Review, Microorganisms, 11(2) doi:10.3390/microorganisms11020510. PMID: 36838475.
3.
Anashkina AA, Petrushanko IY, Ziganshin RH, Orlov YL, Nekrasov AN. (2021), Entropy Analysis of Protein Sequences Reveals a Hierarchical Organization, Entropy (Basel), 23(12) doi:10.3390/e23121647. PMID: 34945953.
4.
Arakawa T. (2018), Protein-solvent interaction, Biophys Rev, 10(2):203-208. doi:10.1007/s12551-017-0339-6. PMID: 29198082.
5.
Berman HM, Westbrook J, Feng Z, Gilliland G, Bhat TN, Weissig H, Shindyalov IN, Bourne PE. (2000), The Protein Data Bank, Nucleic Acids Res, 28(1):235-42. doi:10.1093/nar/28.1.235. PMID: 10592235.
6.
Borrelli GM, Trono D. (2015), Recombinant Lipases and Phospholipases and Their Use as Biocatalysts for Industrial Applications, Int J Mol Sci, 16(9):20774-840. doi:10.3390/ijms160920774. PMID: 26340621.
7.
Burley SK, Berman HM, Duarte JM, Feng Z, Flatt JW, Hudson BP, Lowe R, Peisach E, Piehl DW, Rose Y, Sali A, Sekharan M, Shao C, Vallat B, Voigt M, Westbrook JD, Young JY, Zardecki C. (2022), Protein Data Bank: A Comprehensive Review of 3D Structure Holdings and Worldwide Utilization by Researchers, Educators, and Students, Biomolecules, 12(10) doi:10.3390/biom12101425. PMID: 36291635.
8.
Chakravorty D, Parameswaran S, Dubey VK, Patra S. (2012), Unraveling the rationale behind organic solvent stability of lipases, Appl Biochem Biotechnol, 167(3):439-61. doi:10.1007/s12010-012-9669-9. PMID: 22562495.
9.
Chandra P, Enespa, Singh R, Arora PK. (2020), Microbial lipases and their industrial applications: a comprehensive review, Microb Cell Fact, 19(1):169. doi:10.1186/s12934-020-01428-8. PMID: 32847584.
10.
Chen M, Jin T, Nian B, Cheng W. (2024), Solvent Tolerance Improvement of Lipases Enhanced Their Applications: State of the Art, Molecules, 29(11) doi:10.3390/molecules29112444. PMID: 38893320.
11.
Cheng W, Nian B. (2023), Computer-Aided Lipase Engineering for Improving Their Stability and Activity in the Food Industry: State of the Art, Molecules, 28(15) doi:10.3390/molecules28155848. PMID: 37570817.
12.
Coradetti ST, Pinel D, Geiselman GM, Ito M, Mondo SJ, Reilly MC, Cheng YF, Bauer S, Grigoriev IV, Gladden JM, Simmons BA, Brem RB, Arkin AP, Skerker JM. (2018), Functional genomics of lipid metabolism in the oleaginous yeast Rhodosporidium toruloides, Elife, 7 doi:10.7554/eLife.32110. PMID: 29521624.
13.
Diaz-Villanueva JF, Diaz-Molina R, Garcia-Gonzalez V. (2015), Protein Folding and Mechanisms of Proteostasis, Int J Mol Sci, 16(8):17193-230. doi:10.3390/ijms160817193. PMID: 26225966.
14.
Errami M, Geourjon C, Deleage G. (2003), Conservation of amino acids into multiple alignments involved in pairwise interactions in three-dimensional protein structures, J Bioinform Comput Biol, 1(3):505-520. doi:10.1142/s0219720003000228. PMID: 15307241.
15.
Esque J, Leonard S, de Brevern AG, Oguey C. (2013), VLDP web server: a powerful geometric tool for analysing protein structures in their environment, Nucleic Acids Res, 41(Web Server issue):W373-8. doi:10.1093/nar/gkt509. PMID: 23761450.
16.
Fischer M, Pleiss J. (2003), The Lipase Engineering Database: a navigation and analysis tool for protein families, Nucleic Acids Res, 31(1):319-21. doi:10.1093/nar/gkg015. PMID: 12520012.
17.
Gabanyi MJ, Adams PD, Arnold K, Bordoli L, Carter LG, Flippen-Andersen J, Gifford L, Haas J, Kouranov A, McLaughlin WA, Micallef DI, Minor W, Shah R, Schwede T, Tao YP, Westbrook JD, Zimmerman M, Berman HM. (2011), The Structural Biology Knowledgebase: a portal to protein structures, sequences, functions, and methods, J Struct Funct Genomics, 12(2):45-54. doi:10.1007/s10969-011-9106-2. PMID: 21472436.
18.
Godoy CA, Pardo-Tamayo JS, Barbosa O. (2022), Microbial Lipases and Their Potential in the Production of Pharmaceutical Building Blocks, Int J Mol Sci, 23(17) doi:10.3390/ijms23179933. PMID: 36077332.
19.
Grassmann G, Miotto M, Desantis F, Di Rienzo L, Tartaglia GG, Pastore A, Ruocco G, Monti M, Milanetti E. (2024), Computational Approaches to Predict Protein-Protein Interactions in Crowded Cellular Environments, Chem Rev, 124(7):3932-3977. doi:10.1021/acs.chemrev.3c00550. PMID: 38535831.
20.
Homeyer A, Geissler C, Schwen LO, Zakrzewski F, Evans T, Strohmenger K, Westphal M, Bulow RD, Kargl M, Karjauv A, Munne-Bertran I, Retzlaff CO, Romero-Lopez A, Soltysinski T, Plass M, Carvalho R, Steinbach P, Lan YC, Bouteldja N, Haber D, Rojas-Carulla M, Vafaei Sadr A, Kraft M, Kruger D, Fick R, Lang T, Boor P, Muller H, Hufnagl P, Zerbe N. (2022), Recommendations on compiling test datasets for evaluating artificial intelligence solutions in pathology, Mod Pathol, 35(12):1759-1769. doi:10.1038/s41379-022-01147-y. PMID: 36088478.
21.
Javed S, Azeem F, Hussain S, Rasul I, Siddique MH, Riaz M, Afzal M, Kouser A, Nadeem H. (2018), Bacterial lipases: A review on purification and characterization, Prog Biophys Mol Biol, 132:23-34. doi:10.1016/j.pbiomolbio.2017.07.014. PMID: 28774751.
22.
Kamal MZ, Yedavalli P, Deshmukh MV, Rao NM. (2013), Lipase in aqueous-polar organic solvents: activity, structure, and stability, Protein Sci, 22(7):904-15. doi:10.1002/pro.2271. PMID: 23625694.
23.
Kilgore HR, Raines RT. (2018), n-->pi* Interactions Modulate the Properties of Cysteine Residues and Disulfide Bonds in Proteins, J Am Chem Soc, 140(50):17606-17611. doi:10.1021/jacs.8b09701. PMID: 30403347.
24.
Kouranov A, Xie L, de la Cruz J, Chen L, Westbrook J, Bourne PE, Berman HM. (2006), The RCSB PDB information portal for structural genomics, Nucleic Acids Res, 34(Database issue):D302-5. doi:10.1093/nar/gkj120. PMID: 16381872.
25.
Kumar A, Dhar K, Kanwar SS, Arora PK. (2016), Lipase catalysis in organic solvents: advantages and applications, Biol Proced Online, 18:2. doi:10.1186/s12575-016-0033-2. PMID: 26766927.
26.
Libbrecht MW, Bilmes JA, Noble WS. (2018), Choosing non-redundant representative subsets of protein sequence data sets using submodular optimization, Proteins, 86(4):454-466. doi:10.1002/prot.25461. PMID: 29345009.
27.
Lins L, Thomas A, Brasseur R. (2003), Analysis of accessible surface of residues in proteins, Protein Sci, 12(7):1406-17. doi:10.1110/ps.0304803. PMID: 12824487.
28.
Ma B, Elkayam T, Wolfson H, Nussinov R. (2003), Protein-protein interactions: structurally conserved residues distinguish between binding sites and exposed protein surfaces, Proc Natl Acad Sci U S A, 100(10):5772-7. doi:10.1073/pnas.1030237100. PMID: 12730379.
29.
Masica DL, Ash JT, Ndao M, Drobny GP, Gray JJ. (2010), Toward a structure determination method for biomineral-associated protein using combined solid- state NMR and computational structure prediction, Structure, 18(12):1678-87. doi:10.1016/j.str.2010.09.013. PMID: 21134646.
30.
Masson P, Lushchekina S. (2022), Conformational Stability and Denaturation Processes of Proteins Investigated by Electrophoresis under Extreme Conditions, Molecules, 27(20) doi:10.3390/molecules27206861. PMID: 36296453.
31.
Mieres-Perez J, Almeida-Hernandez Y, Sander W, Sanchez-Garcia E. (2025), A Computational Perspective to Intermolecular Interactions and the Role of the Solvent on Regulating Protein Properties, Chem Rev, 125(15):7023-7056. doi:10.1021/acs.chemrev.4c00807. PMID: 40720534.
32.
Niazi SK. (2025), Protein Catalysis Through Structural Dynamics: A Comprehensive Analysis of Energy Conversion in Enzymatic Systems and Its Computational Limitations, Pharmaceuticals (Basel), 18(7) doi:10.3390/ph18070951. PMID: 40732241.
33.
Pace CN, Trevino S, Prabhakaran E, Scholtz JM. (2004), Protein structure, stability and solubility in water and other solvents, Philos Trans R Soc Lond B Biol Sci, 359(1448):1225-34; discussion 1234-5. doi:10.1098/rstb.2004.1500. PMID: 15306378.
34.
Pan Y, Zhou S, Guan J. (2020), Computationally identifying hot spots in protein-DNA binding interfaces using an ensemble approach, BMC Bioinformatics, 21(Suppl 13):384. doi:10.1186/s12859-020-03675-3. PMID: 32938375.
35.
Panja AS, Maiti S, Bandyopadhyay B. (2020), Protein stability governed by its structural plasticity is inferred by physicochemical factors and salt bridges, Sci Rep, 10(1):1822. doi:10.1038/s41598-020-58825-7. PMID: 32020026.
36.
Pilla KB, Otting G, Huber T. (2017), 3D Computational Modeling of Proteins Using Sparse Paramagnetic NMR Data, Methods Mol Biol, 1526:3-21. doi:10.1007/978-1-4939-6613-4_1. PMID: 27896733.
37.
Qing R, Hao S, Smorodina E, Jin D, Zalevsky A, Zhang S. (2022), Protein Design: From the Aspect of Water Solubility and Stability, Chem Rev, 122(18):14085-14179. doi:10.1021/acs.chemrev.1c00757. PMID: 35921495.
38.
Rosgen J, Pettitt BM, Bolen DW. (2005), Protein folding, stability, and solvation structure in osmolyte solutions, Biophys J, 89(5):2988-97. doi:10.1529/biophysj.105.067330. PMID: 16113118.
39.
Safari-Alighiarloo N, Taghizadeh M, Rezaei-Tavirani M, Goliaei B, Peyvandi AA. (2014), Protein-protein interaction networks (PPI) and complex diseases, Gastroenterol Hepatol Bed Bench, 7(1):17-31. doi: https://www.ncbi.nlm.nih.gov/p.... PMID: 25436094.
40.
Savojardo C, Manfredi M, Martelli PL, Casadio R. (2020), Solvent Accessibility of Residues Undergoing Pathogenic Variations in Humans: From Protein Structures to Protein Sequences, Front Mol Biosci, 7:626363. doi:10.3389/fmolb.2020.626363. PMID: 33490109.
41.
Sharma S, Kanwar SS. (2014), Organic solvent tolerant lipases and applications, ScientificWorldJournal, 2014:625258. doi:10.1155/2014/625258. PMID: 24672342.
42.
Singh P, Ramanathan V, Sankararamakrishnan R. (2024), Self-contacting Cys, Ser, and Thr residues in high-resolution protein crystal structures: Tertiary constraints or hydrogen bonds?, Protein Sci, 33(12):e5218. doi:10.1002/pro.5218. PMID: 39548825.
43.
Soni S. (2022), Trends in lipase engineering for enhanced biocatalysis, Biotechnol Appl Biochem, 69(1):265-272. doi:10.1002/bab.2105. PMID: 33438779.
44.
Sraphet S, Javadi B. (2022), Application of Hierarchical Clustering to Analyze Solvent-Accessible Surface Area Patterns in Amycolatopsis lipases, Biology (Basel), 11(5) doi:10.3390/biology11050652. PMID: 35625380.
45.
Standley DM, Kinjo AR, Kinoshita K, Nakamura H. (2008), Protein structure databases with new web services for structural biology and biomedical research, Brief Bioinform, 9(4):276-85. doi:10.1093/bib/bbn015. PMID: 18430752.
46.
Szklarczyk D, Gable AL, Nastou KC, Lyon D, Kirsch R, Pyysalo S, Doncheva NT, Legeay M, Fang T, Bork P, Jensen LJ, von Mering C. (2021), The STRING database in 2021: customizable protein-protein networks, and functional characterization of user-uploaded gene/measurement sets, Nucleic Acids Res, 49(D1):D605-D612. doi:10.1093/nar/gkaa1074. PMID: 33237311.
47.
Szymczyna BR, Taurog RE, Young MJ, Snyder JC, Johnson JE, Williamson JR. (2009), Synergy of NMR, computation, and X-ray crystallography for structural biology, Structure, 17(4):499-507. doi:10.1016/j.str.2009.03.001. PMID: 19368883.
48.
Tan KP, Nguyen TB, Patel S, Varadarajan R, Madhusudhan MS. (2013), Depth: a web server to compute depth, cavity sizes, detect potential small-molecule ligand-binding cavities and predict the pKa of ionizable residues in proteins, Nucleic Acids Res, 41(Web Server issue):W314-21. doi:10.1093/nar/gkt503. PMID: 23766289.
49.
Tan KP, Varadarajan R, Madhusudhan MS. (2011), DEPTH: a web server to compute depth and predict small-molecule binding cavities in proteins, Nucleic Acids Res, 39(Web Server issue):W242-8. doi:10.1093/nar/gkr356. PMID: 21576233.
50.
Thompson R, Pickard BS. (2024), The amino acid composition of a protein influences its expression, PLoS One, 19(10):e0284234. doi:10.1371/journal.pone.0284234. PMID: 39401228.
51.
Tien MZ, Meyer AG, Sydykova DK, Spielman SJ, Wilke CO. (2013), Maximum allowed solvent accessibilites of residues in proteins, PLoS One, 8(11):e80635. doi:10.1371/journal.pone.0080635. PMID: 24278298.
52.
Vardar-Yel N, Tutuncu HE, Surmeli Y. (2024), Lipases for targeted industrial applications, focusing on the development of biotechnologically significant aspects: A comprehensive review of recent trends in protein engineering, Int J Biol Macromol, 273(Pt 1):132853. doi:10.1016/j.ijbiomac.2024.132853. PMID: 38838897.
53.
Wlodawer A, Minor W, Dauter Z, Jaskolski M. (2008), Protein crystallography for non-crystallographers, or how to get the best (but not more) from published macromolecular structures, FEBS J, 275(1):1-21. doi:10.1111/j.1742-4658.2007.06178.x. PMID: 18034855.
54.
Zhou H, Zhou Y. (2004), Quantifying the effect of burial of amino acid residues on protein stability, Proteins, 54(2):315-22. doi:10.1002/prot.10584. PMID: 14696193.
We process personal data collected when visiting the website. The function of obtaining information about users and their behavior is carried out by voluntarily entered information in forms and saving cookies in end devices. Data, including cookies, are used to provide services, improve the user experience and to analyze the traffic in accordance with the Privacy policy. Data are also collected and processed by Google Analytics tool (more).
You can change cookies settings in your browser. Restricted use of cookies in the browser configuration may affect some functionalities of the website.
You can change cookies settings in your browser. Restricted use of cookies in the browser configuration may affect some functionalities of the website.